Why openly available abstracts are important — overview of the current state of affairs
Openness of the metadata of scientific articles is increasingly being discussed. In this blog post, Aaron Tay (SMU Libraries, Singapore Management University), Bianca Kramer (Utrecht University Library), and Ludo Waltman (CWTS, Leiden University) discuss the value of openly available abstracts.
This post was originally published at Medium.
The value of open and interoperable metadata of scientific articles is increasingly being recognized, as demonstrated by the work of organizations such as Crossref, DataCite, and OpenCitations and by initiatives such as Metadata 2020 and the Initiative for Open Citations. At the same time, scientific articles are increasingly being made openly accessible, stimulated for instance by Plan S, AmeliCA, and recent developments in the US, and also by the need for open access to coronavirus literature.
In this post, we focus on a key issue at the interface of these two developments: The open availability of abstracts of scientific articles. Abstracts provide a summary of an article and are part of an article’s metadata. We first discuss the many ways in which abstracts can be used and we then explore the availability of abstracts. The open availability of abstracts is surprisingly limited. This creates important obstacles to scientific literature search, bibliometric analysis, and automatic knowledge extraction.
The many uses of abstracts
The most basic way in which researchers benefit from abstracts is clear: Abstracts provide a summary of an article and are used by researchers to quickly decide whether an article is likely to be of interest. In addition, however, abstracts are also used in a large number of more advanced ways.
Abstracts of course play an important role in scientific literature search. Databases such as Web of Science, Scopus, and PubMed enable searching in abstracts of articles, and so do many lesser known alternatives (e.g., Scilit and Open Ukrainian Citation Index). Other databases (e.g., Google Scholar) support full-text search, but in many cases (e.g., Dimensions, Lens, and PubMed Central) these databases also offer the possibility to restrict a search to abstracts only, which will give more focused results.

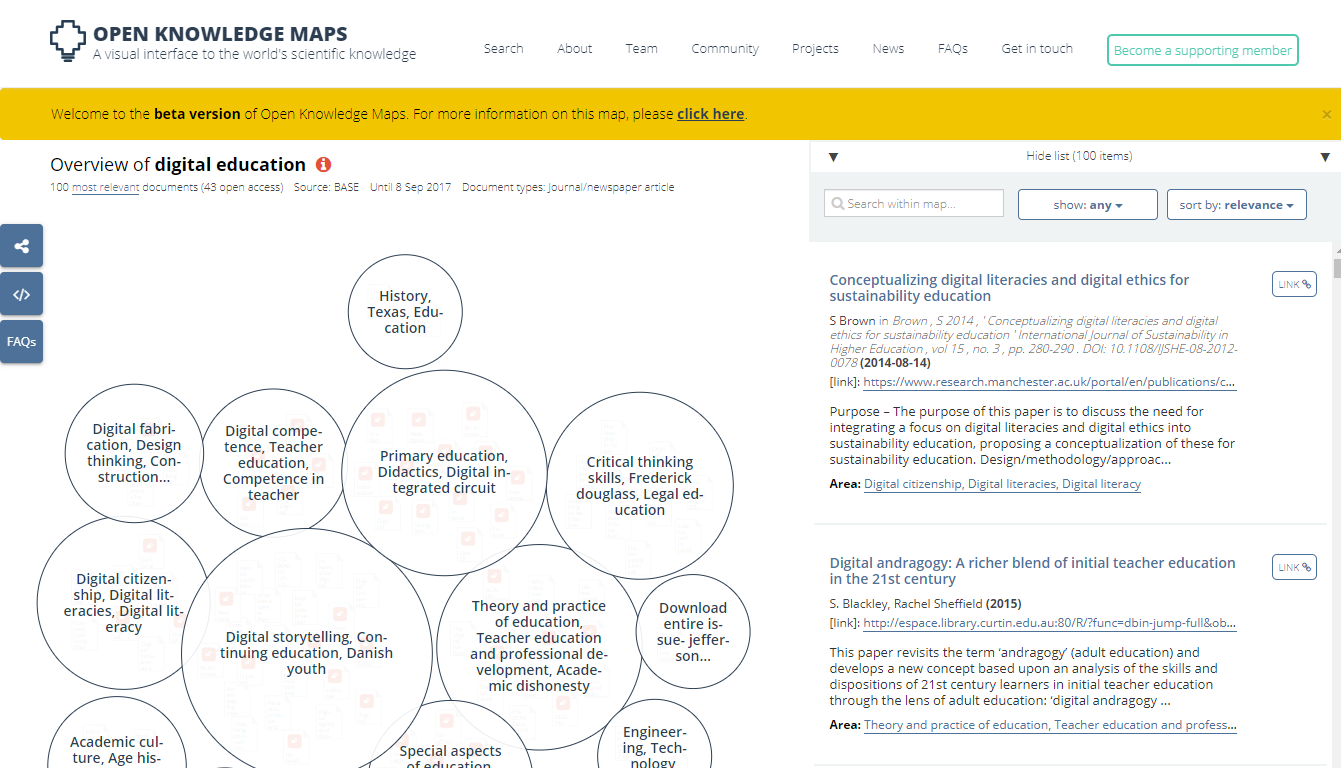
Discovery tools increasingly make use of text mining features combined with visual user interfaces. Open Knowledge Maps is a prominent example. Iris.ai is another one. These tools group articles based on textual similarity of titles and abstracts and visually show the resulting clusters of articles.
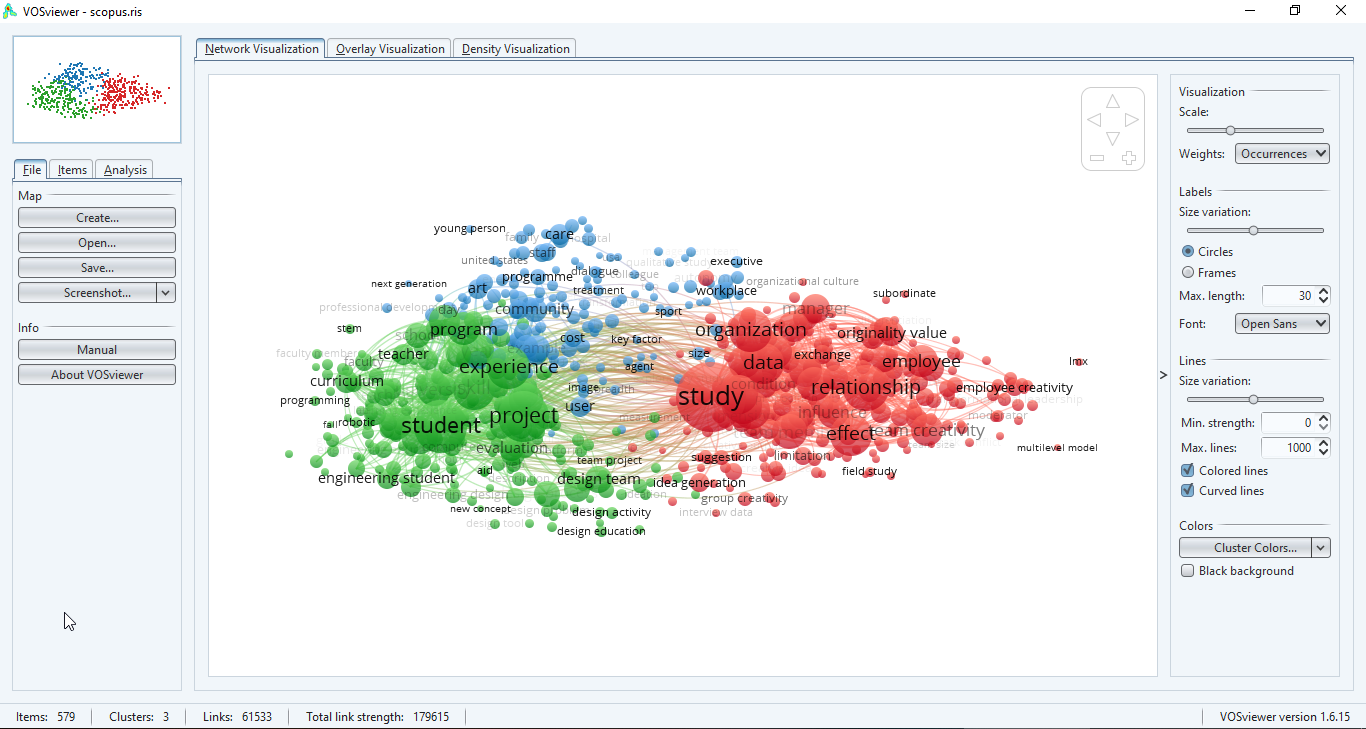
Bibliometric visualization tools such as VOSviewer and CiteSpace perform text mining of titles and abstracts of articles to offer an overview of the literature in a research field. VOSviewer uses text mining to create term co-occurrence maps that show the main topics studied in a field. CiteSpace analyzes emerging trends and research fronts using text mining.
Abstracts can also be used to improve the functionality of many library systems, such as library discovery systems like Primo and Summon, research information systems like Pure and Converis, and more. Institutional repository (IR) managers can improve the discoverability of their IR content by enhancing IR entries with abstracts, similar to what CORE has done.
Some libraries, for instance in Finland, use automated methods for subject indexing of their IR content. This can be done based on titles and abstracts of articles. It doesn’t require full texts. Microsoft Academic for instance uses abstracts, not full texts, for tagging fields of study to articles.
A different way of using abstracts is illustrated by Get The Research, which aims to help readers, in particular from outside academia, find and understand scientific articles. Titles and abstracts of articles, taken from PubMed, are enriched by adding links to plain language explanations of scientific terms, obtained from Wikipedia. In this case, abstracts have a benefit for a wide audience, also outside academia.
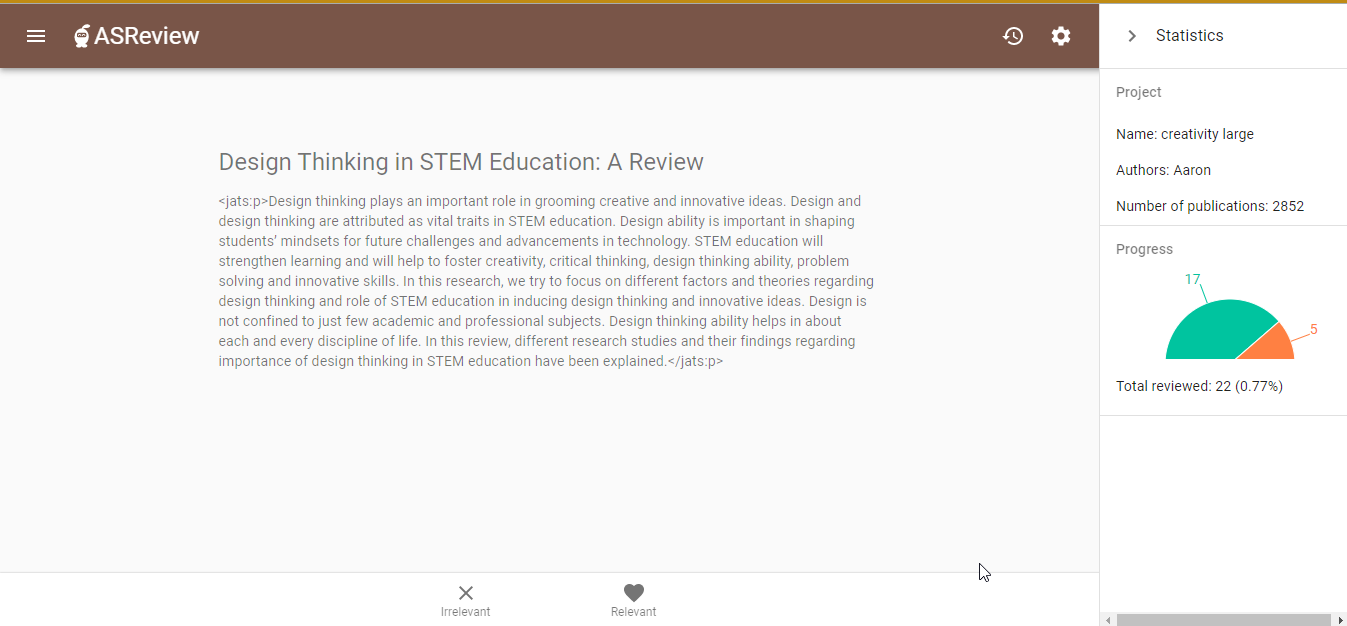
Abstracts also play a crucial role in systematic reviewing. Literature searches for systematic reviews are usually done in the titles, abstracts, and keywords of articles, and the search results are typically screened (manually and sometimes assisted by machine learning) based on titles and abstracts. Hence, abstracts are essential for high-quality systematic reviewing. An example is the screening of the CORD-19 dataset of COVID-19 related articles using the ASReview tool. However, the fact that abstracts are missing for 22% of the articles in the full CORD-19 dataset, and even for 36% of the CORD-19 articles from 2020, limits the use of the dataset.
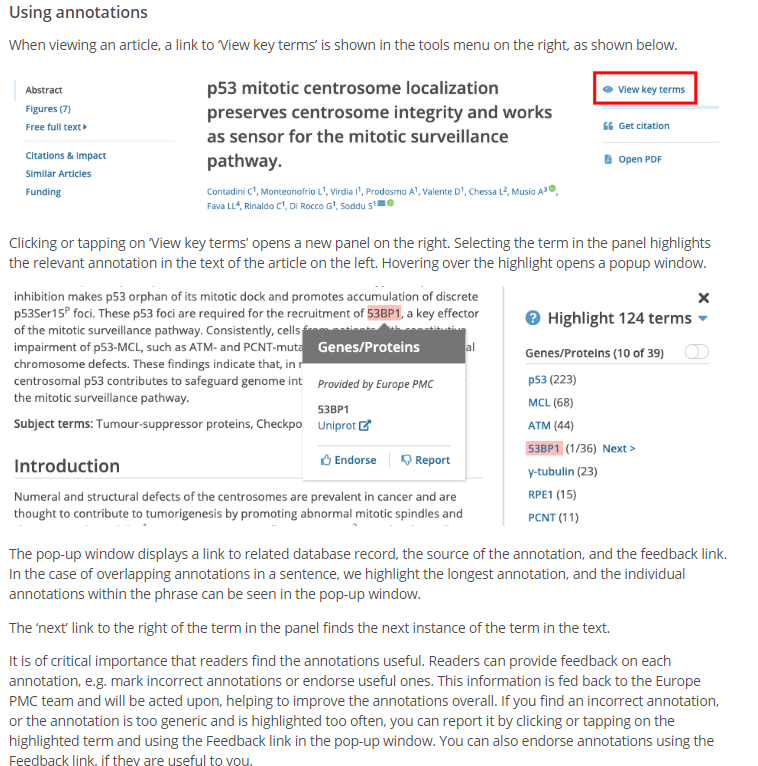
The most sophisticated uses of abstracts arguably can be found in the biomedical domain, where the open availability of abstracts in PubMed has spurred innovation. One example is PubMed’s feature for finding similar articles, which compares articles based on their titles and abstracts. Another example can be found in Europe PMC, where the SciLite platform provides annotations of biomedical entities identified in abstracts (and full text, if available) using text mining algorithms. Identification of biomedical entities in abstracts enables computational analyses of associations between these entities. Gene-disease associations have for instance been extracted from abstracts by the LHGDN (Literature-derived human gene-disease network) and BeFree systems and have been made available in the DisGeNET database. In a similar way, researchers have developed algorithms for extracting protein-protein interactions from the abstracts of biomedical articles. Although similar approaches to the use of abstracts for automatic knowledge extraction are also being explored in other fields of research (e.g., materials science), the biomedical domain is clearly ahead of these other fields. This is most likely due to the open availability of abstracts in PubMed.
As the above examples illustrate, abstracts have many uses. This is true even given the increasing number of articles for which the full text is openly accessible. While it would often be beneficial to have access to full texts rather than just abstracts, such access is unfortunately not yet a given for the full scholarly literature. In many cases, however, it is a deliberate choice to work with abstracts rather than full texts, even when the full text of articles is accessible. This can be for technical reasons, but also because abstracts provide a more focused description of the underlying research. For instance, a recent analysis of the CORD-19 dataset of COVID-related articles showed that only 24% of the CORD-19 articles available in Web of Science include COVID-related terms in their title, abstract, or keywords — restricting to this subset may give more targeted results than using the full dataset.
Availability of abstracts
The ways in which researchers benefit from abstracts are quite varied. But are abstracts generally available for the various use cases discussed above?
Abstracts are mostly freely accessible on publisher websites. This facilitates the most basic use case for abstracts, namely to enable researchers to easily get a general understanding of what an article is about. However, for the other use cases discussed above, having free access to abstracts on publisher websites is not enough. These use cases require bulk access to machine readable abstracts.
Commercial and proprietary bibliographic databases such as Web of Science and Scopus provide a good coverage of abstracts. However, in addition to the cost of accessing them, they have the limitation of being selective in scope, not covering the full breadth of the scholarly literature. Moreover, they impose restrictions on the number of abstracts that can be downloaded from the database and the way these abstracts can be used. Dimensions is generally less selective and provides free access, but it currently has the disadvantage that abstracts cannot be downloaded. (The Dimensions team informed us that they plan to make abstracts available for download in the near future.)
Microsoft Academic, which is openly available under an ODC-BY license, provides some hope though. It is pretty comprehensive in scope and is also being used by other databases such as the Lens and Semantic Scholar. However, as Microsoft Academic obtains its data by scraping the World Wide Web, abstracts suffer from some quality problems. In an analysis of abstracts in Microsoft Academic, we for instance found that abstracts are sometimes truncated and that they sometimes include text that actually doesn’t belong to the abstract (e.g., author and affiliation data or the opening paragraph of an article).
Like Microsoft Academic, PubMed makes abstracts openly available. Since PubMed receives abstracts directly from publishers, the data quality is higher than in the case of Microsoft Academic. However, PubMed has the limitation of being restricted to biomedical research.
Another cross-domain source of bibliographic metadata, including abstracts, is CORE (COnnecting REpositories) — one of the largest aggregators of research articles harvested from subject and institutional repositories as well as open access and hybrid journals. The data is free to access and for non-commercial purposes the data can also be freely downloaded. CORE content is for instance used in the Lens. The exact coverage of CORE, especially for metadata of paywalled articles, is hard to ascertain.
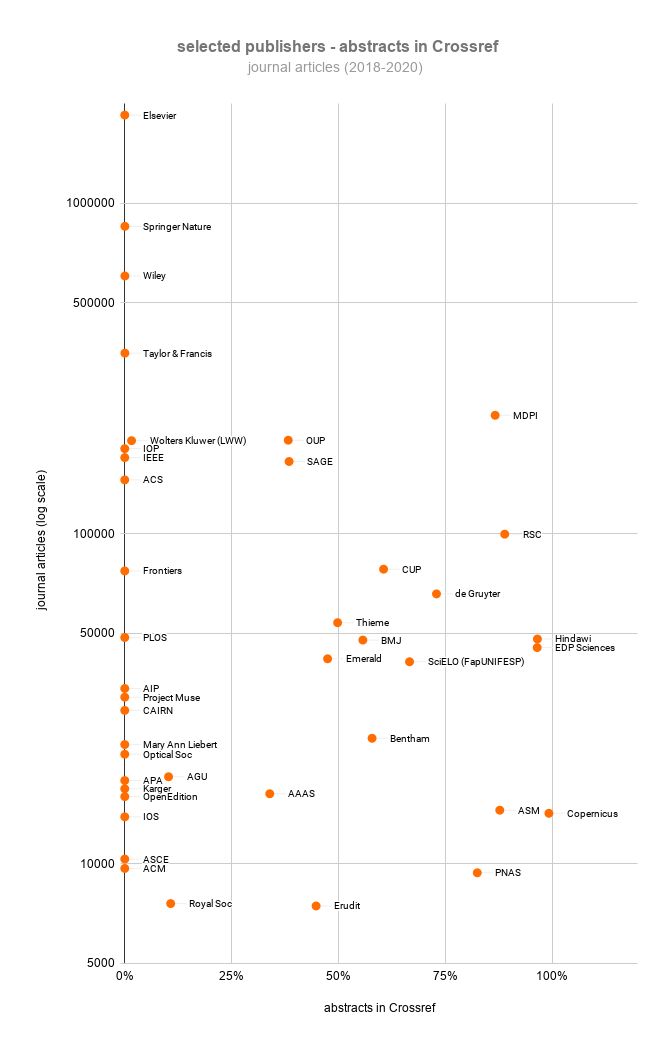
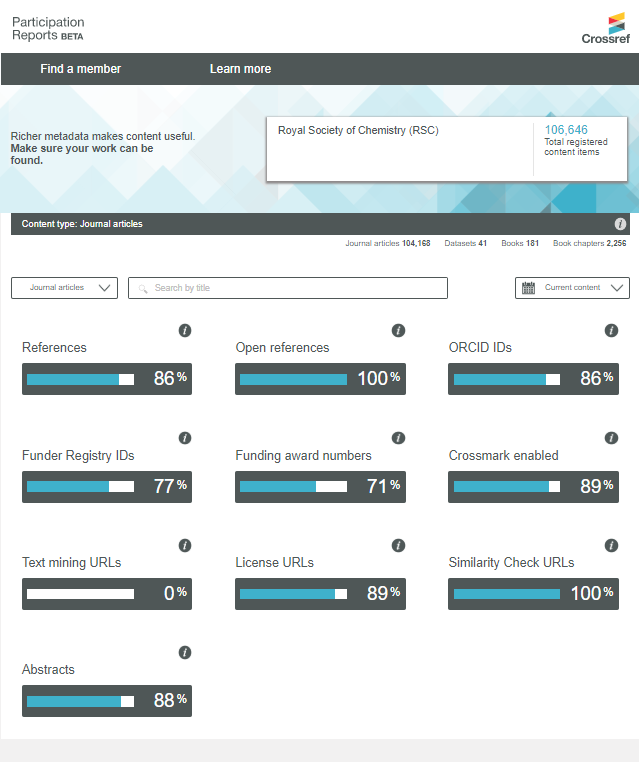
Finally, an important source of bibliographic metadata is provided by Crossref, a not-for-profit organization that most international publishers work with to register Digital Object Identifiers (DOIs) for their content. Except for some citation data, all metadata in Crossref is openly available. Unlike Microsoft Academic, Crossref receives its metadata directly from publishers, resulting in a higher data quality, and unlike PubMed, Crossref is not restricted to biomedical research. However, although Crossref offers a first-class infrastructure for making abstracts openly available, the actual number of abstracts in Crossref is disappointingly low. At the moment, only 8% of all journal articles in Crossref have an abstract. For recent years, this percentage is somewhat higher (20% for 2018–2020). As shown in the figure above, many publishers do not make abstracts available in Crossref, or they have done so only for a small share of their articles. More detailed information for individual publishers can be found in Crossref’s Participation Reports.
Making abstracts openly available
As we have seen, abstracts are of great value for scientific literature search, bibliometric analysis, and automatic knowledge extraction. Innovative uses of abstracts can be found in particular in biomedical fields, which benefit from the open availability of abstracts in PubMed. However, outside the biomedical domain, the lack of a centralized database in which abstracts are made openly available hinders the development of innovative new tools to support researchers. This will inevitably slow down the speed at which researchers come up with new ideas and make new discoveries.
Microsoft Academic is the most comprehensive open source of abstracts. It is of great value, but it has two limitations. First, since abstracts have been scraped from the Web, they suffer from data quality issues. And second, we don’t know much about the long-term prospects of Microsoft Academic, which depend on Microsoft’s willingness to continue investing in Microsoft Academic and to keep the data open.
Crossref clearly provides the most promising centralized infrastructure for making abstracts openly available. While this infrastructure is readily available to all publishers working with Crossref, many of them unfortunately do not yet make use of it, perhaps because they are not aware of it or do not see its value.
Thanks to the Initiative for Open Citations, a large number of publishers have made the reference lists of their articles openly available in Crossref. In the same spirit, we call on publishers to make abstracts openly available in Crossref. Publishers already make the abstracts of articles in biomedical fields openly available in PubMed, so why not use the infrastructure of Crossref to do the same for articles in all fields? By making this small effort, publishers not only give additional visibility to their content, but they also make a large contribution to the benefit of science.
This post is licensed under a CC BY 4.0 license.





0 Comments
Add a comment