Delineating COVID-19 and coronavirus research
Many initiatives are keeping track of research on COVID-19 and coronaviruses. These initiatives, while valuable because they allow for fast access to relevant research, pose the question of subject delineation. We analyse here one such initiative, the COVID-19 Open Research Dataset (CORD-19).
As the COVID-19 pandemic unfolds, researchers from all disciplines are coming together to contribute their expertise. The COVID-19 Open Research Dataset (CORD-19) is a growing, weekly-updated dataset of COVID-19 publications, capturing new as well as past research on “COVID-19 and the coronavirus family of viruses for use by the global research community.” The reason to release this dataset is “to mobilize researchers to apply recent advances in natural language processing to generate new insights in support of the fight against this infectious disease.” The initiative is a partnership of several institutions including the Chan Zuckerberg Initiative, Georgetown University's Center for Security and Emerging Technology, Microsoft Research, the National Library of Medicine of the National Institutes of Health, and Unpaywall. CORD-19 is released together with a set of challenges hosted by Kaggle, mainly focused on automatically extracting structured and actionable information from such a large set of publications. The release of CORD-19 is a call for action directed towards the natural language processing, machine learning, and related research communities. This call has been taken up. For example, the ACL conference has announced an emergency NLP COVID-19 workshop, and a TREC-COVID challenge has been released, both using CORD-19.
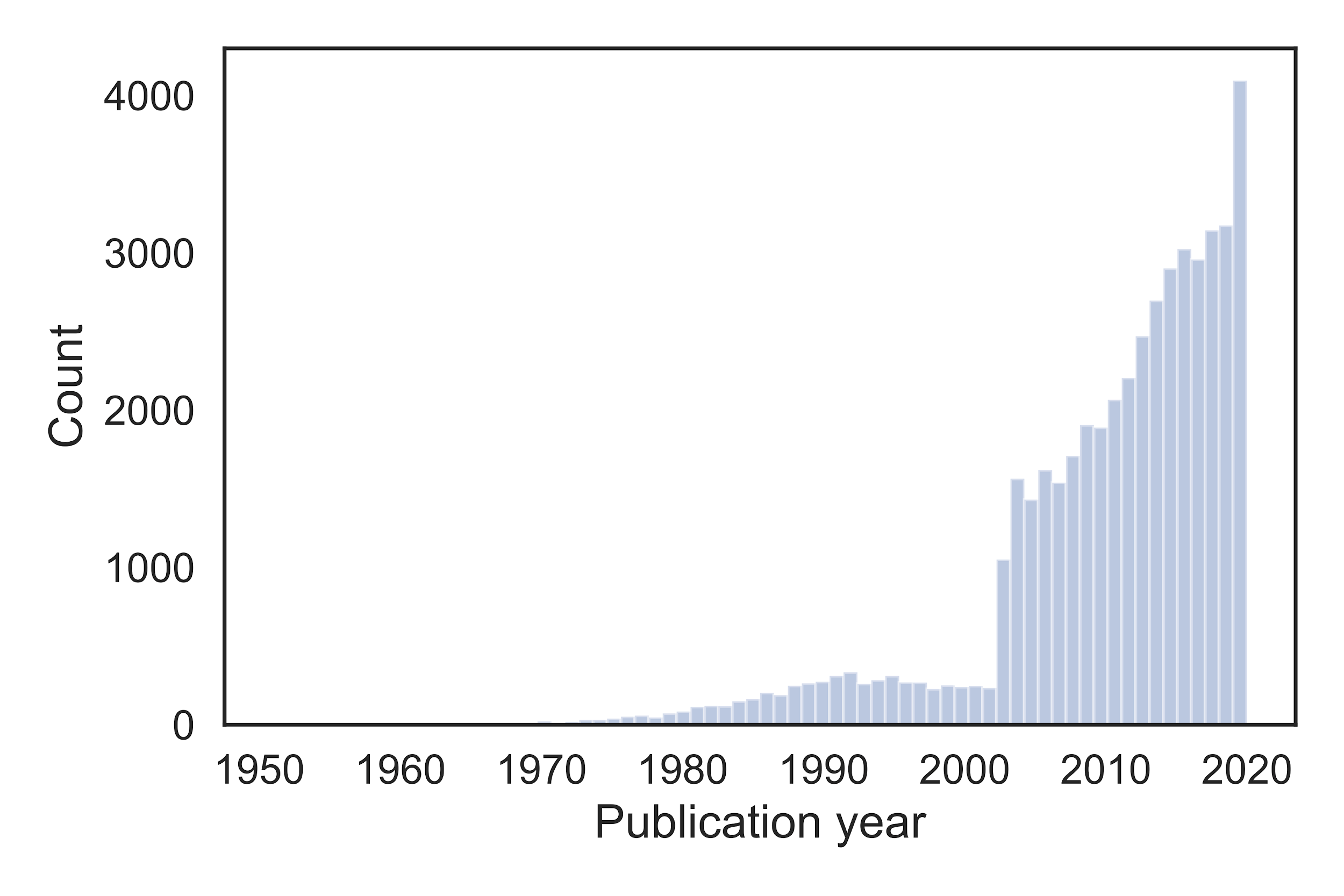
CORD-19 contains over 47,000 articles, of which about 36,000 are equipped with full text, as recorded on April 4, 2020. Before the SARS outbreak in 2003, there were only few publications on the subject. The number of publications steadily increased in the years afterwards. In 2020 it reaches a peak, with thousands of publications in the first few months alone.
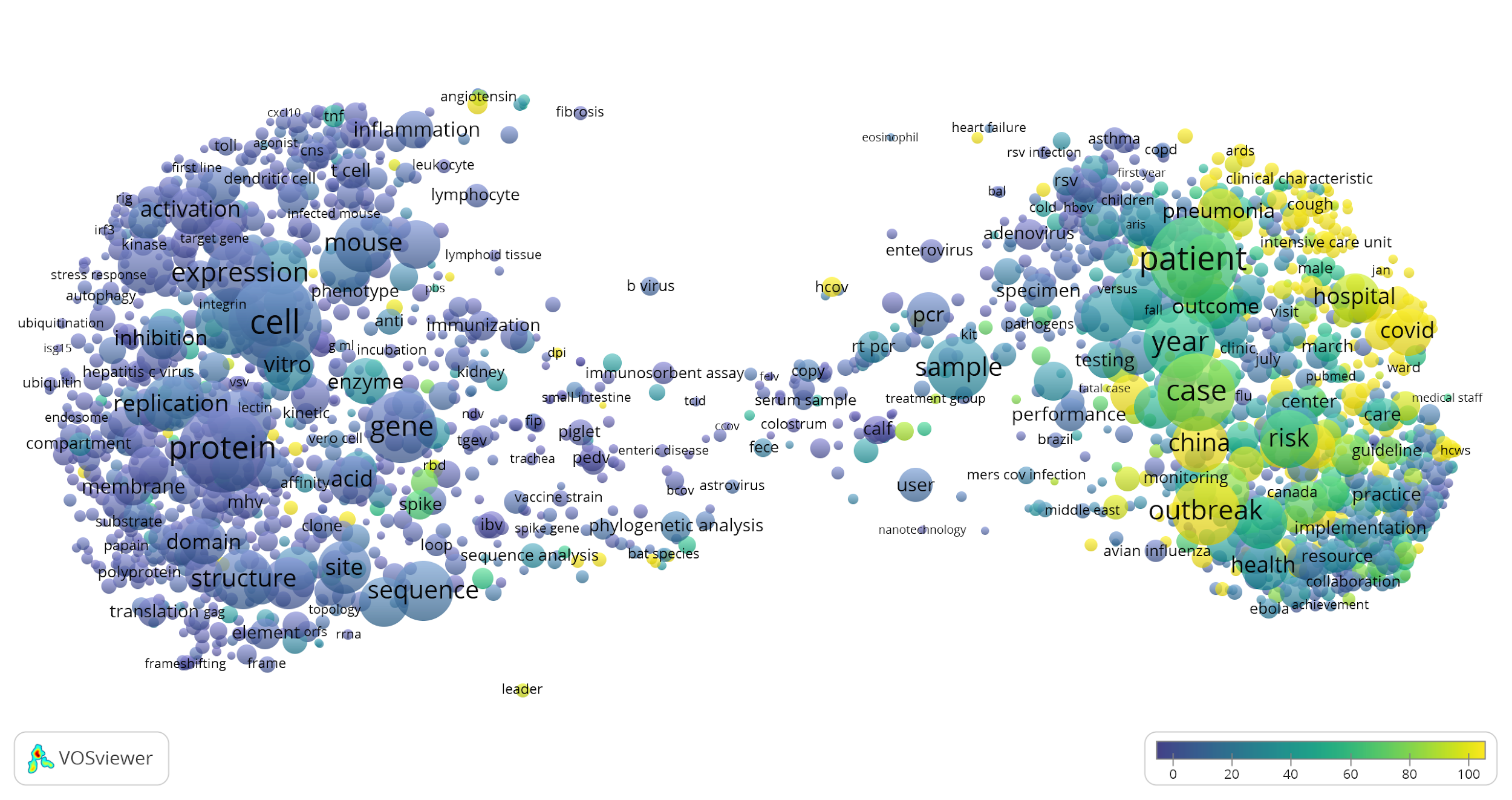
To get a high-level overview of the CORD-19 dataset, we used VOSviewer to create a term map of the publications in this dataset. The size of a term reflects the number of publications in which the term occurs. The proximity of two terms in the map indicates how strongly related the two terms are, based on how frequently they co-occur. The closer two terms are located to each other, the stronger they are related. The map shows a clear divide between biologically focused topics in the left part of the map and clinically focused topics and health research in the right part. In the visualization of the map, presented below, the colour of a term reflects the average publication year of the publications in which the term occurs.
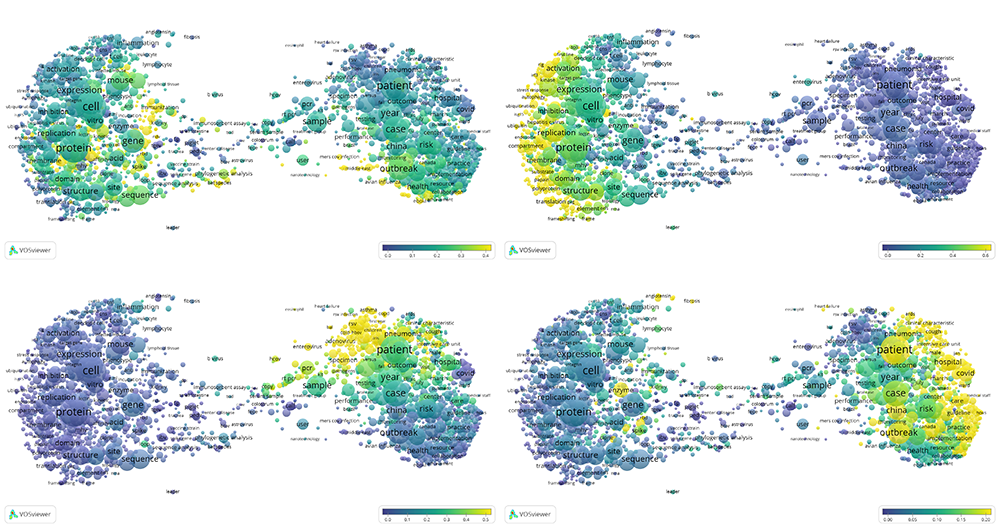
Four large clusters of the CORD-19 citation network (see also the interactive term map).
Using topic modeling we further characterised these four largest clusters. The largest cluster (top-left) mainly contains publications on coronaviruses and their molecular biology. The second-largest cluster (top-right) focuses on molecular biology and immunology. The third-largest cluster (bottom-left) represents research on influenza and related viruses. Finally, the fourth-largest cluster (bottom-right) contains publications on coronavirus outbreaks, their clinical features and epidemiological impact. These areas of research are interrelated, yet also contain specialised information, highlighting distinct research topics within CORD-19: coronaviruses, molecular biology research on viruses, public health and epidemics, other viruses (such as influenza) and other related topics (immunology, diagnosing, trials and testing).
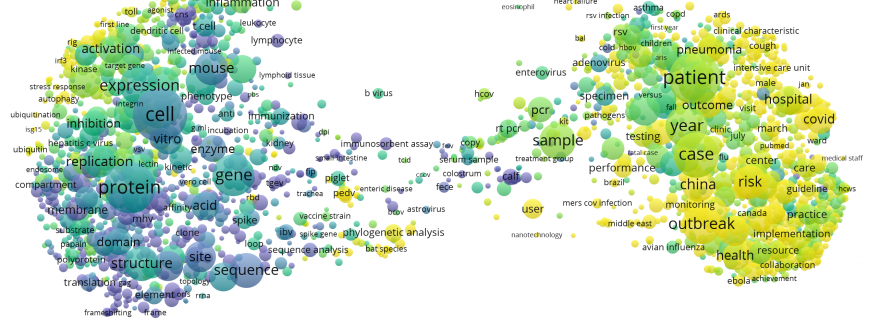
We also analysed CORD-19 publications using Altmetric data, with the aim of exploring the reception of these publications on social media. Overall 60% of the publications in CORD-19 have received some mention in Altmetric, and for publications from 2020, this is the case for more than 80%. Twitter is the leading Altmetric source for CORD-19 publications, especially in 2020. As shown below, CORD-19 publications mentioned on Twitter can be found mainly in the right part of the term map, focused mostly on the epidemics and clinical characteristics of the current COVID-19 outbreak.
Twitter attention of CORD-19 publications (see also the interactive term map).
Our most important finding is that CORD-19 contains research not only on COVID-19 and coronaviruses but on viruses in general. In fact, only approximately 11,500 of the 47,000 CORD-19 publications include coronavirus-related terms in their title or abstract. While we fully endorse the initiative that led to CORD-19, it is important to be aware of the relatively broad content of the dataset. In addition, there also seems to be research on coronaviruses that is missing in CORD-19. We were able to identify almost 5,000 publications in the Web of Science that are not included in CORD-19 even though they explicitly mention coronaviruses or related terms. Many of these publications can even be found in PubMed, but are nonetheless not included in the CORD-19 dataset.
In this blog post, we just started to scratch the surface of the potential of the CORD-19 dataset. There are many open scientometric challenges on this dataset, and on COVID-19 and coronavirus research more broadly. For example, there is a need for a more comprehensive and multidisciplinary map of COVID-19-related research, going beyond biomedical research. CORD-19 also provides a virtuous example of open data sharing, and the scientometric community can contribute by creating and maintaining additional datasets on COVID-19 research. Furthermore, we showed that there is a lot of social media attention for COVID-19 research, which calls for more advanced analytics on the reception of this research in social media . Understanding the mechanics behind the online dissemination of COVID-19 research can inform science communication strategies, and provide valuable advice to experts and governments during the current and future pandemics.







0 Comments
Add a comment